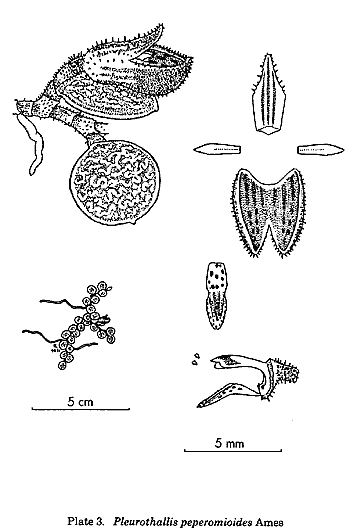
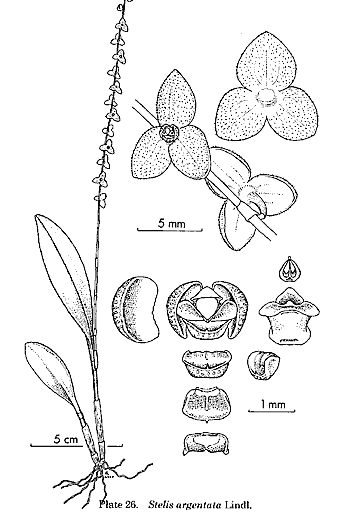
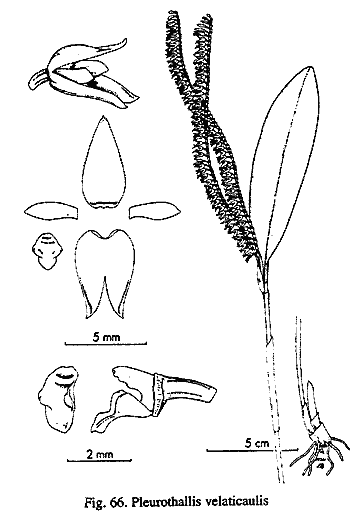
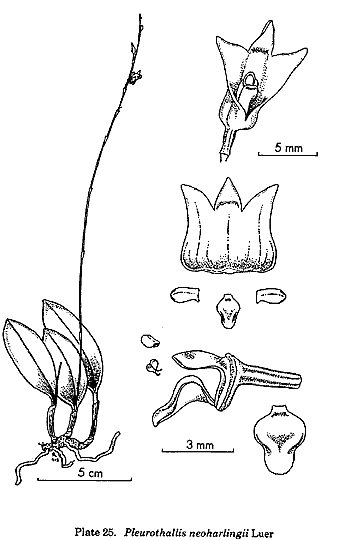
By
Lou Jost and Lorena Endara
Note added Jan 1, 2006: In the last two months many new results have been announced by Alec Pridgeon and colleagues, by Dr. Luer, by Hagen Stenzel, among others. These results help clear up some of the problem areas we have identified in this article. Hopefully we will soon find the time to incorporate these new results into the present article.
Like everyone else in the Pleurothallid world, we are trying to figure out what to do with Pridgeon, Solano, and Chases proposed reclassification of our favorite subtribe based on DNA analysis. We are trying to come to grips with this by looking closely at the internal logic of the proposal. We think that the discussion so far has oversimplified the issue. Many of their results mirror Dr Luers morphology-based taxonomy, so we cannot simply say that the whole DNA analysis is garbage. On the other hand, some of their results deviate so greatly from the morphological evidence that they invite skepticism. We must look carefully at the details of each of the changes proposed by Pridgeon and his collaborators, accepting those that are well-supported and holding off on the poorly supported ones. We feel strongly that the correct approach is not simply to accept or reject either this reclassification or the traditional one. We need all the clues we can get to help us with the taxonomy of this difficult group, and it would be a shame to just throw away either Pridgeons work or Dr Luers lifetime of insights, as some have done. In spite of Pridgeon et als rhetoric
against a morphology-based taxonomy, we are impressed with the
overall convergence between Dr Luers morphology-based taxonomy
and the DNA-based taxonomy of Pridgeon and company. Dr. Luers
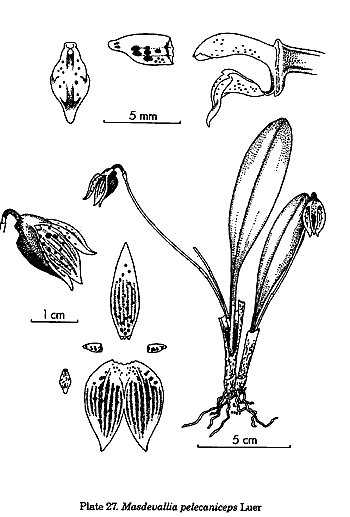
splitting of Dracula, Trisetella, and Dryadella
from Masdevallia, his splitting of Trichosalpinx,
Zootrophion, Dresslerella, Frondaria,
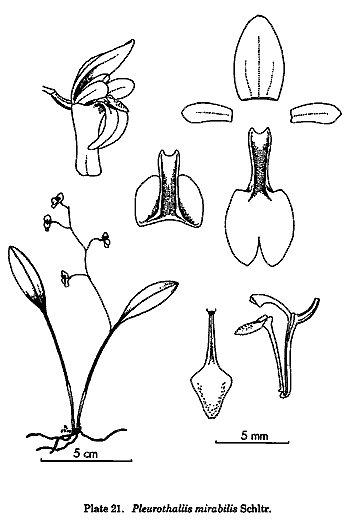
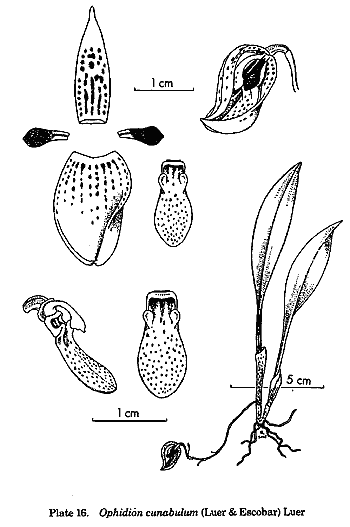
and Ophidion from Pleurothallis, and his insight
that Salpistele consists of two unrelated groups (first
mentioned in Luer 1991), are all strongly supported by the work
of Pridgeon et al. In general, Dr Luers major subgeneric classifications
are also congruent with the
DNA-based groupings, so much so that Pridgeon et al rely heavily
on Dr Luers groupings to make nomenclatural transfers of species they have not tested. Dr Luer criticizes
this, but we see it rather as a tacit homage to the accuracy
of Dr Luers work. The major difference between the two classifications
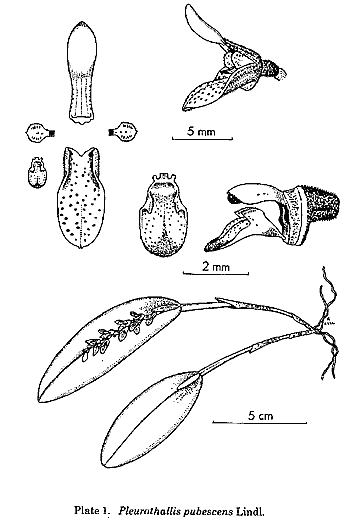
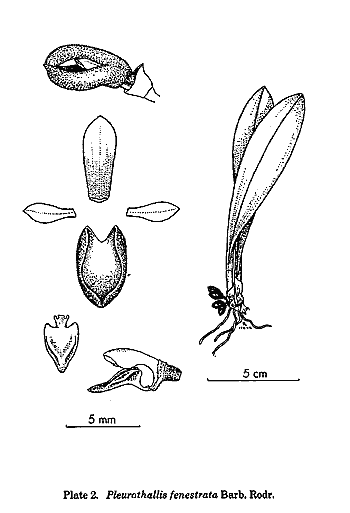
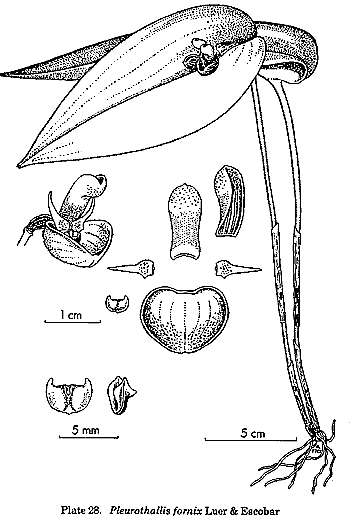
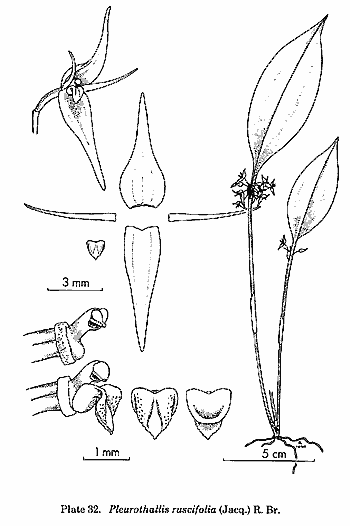
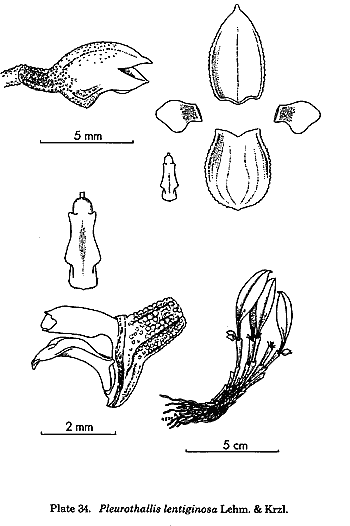
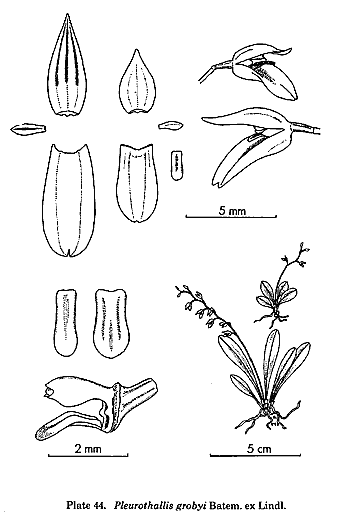
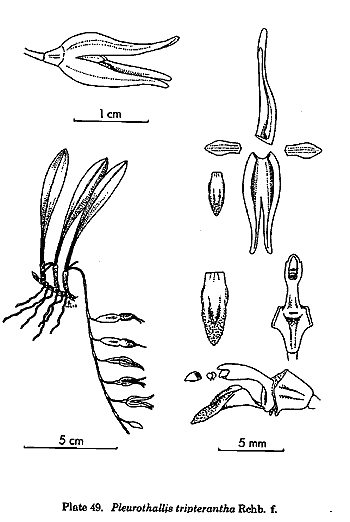
is the treatment of the traditional genus Pleurothallis,
but this genus had always been a dumping-ground for species
with no distinctive features that could lift them into other
genera, and it should come as no surprise that it can be broken
up with the help of new genetic clues. |